MIB2 is a binding sites prediction and modeling server for metal ions, and this server provides an accurate, integrated method to search the residues in metal ion-binding sites by using the fragment transformation method. MIB2 utilizes both the (PS)2 method and the AlphaFold protein structure database to acquire 3D information to perform metal ion docking and predict binding residues. As an updated version of MIB, MIB2 offers a total of 18 types of metal ions (Ca2+, Cu2+, Fe3+, Mg2+, Mn2+, Zn2+, Cd2+, Fe2+ ,Ni2+, Hg2+, Co2+, Cu+, Au+, Ba2+, Pb2+, Pt2+, Sm3+, and Sr2+), supportive of binding site predictions. A reliable protein structure model can be built with a total of 18 metal ion types.
Recently, there are more than 180,000 structures in Protein Data Bank (PDB), and a large amount of proteins, approximately one-third contain metal ions essential for their functions. In MIB2, we utilize both the (PS)2 method and the AlphaFold protein structure database to acquire 3D structure information for those proteins without structures in PDB. Metal ions found in proteins include those of the alkali metals, alkaline earth metals and transition metals, with the most common being sodium and potassium ions, calcium and magnesium ions, and iron, manganese, copper, and zinc ions, respectively. For the metal ion–binding proteins found in the Protein Data Bank, ~66 % contain transition metal ions, ~37 % contain alkaline earth metal ions, and ~6 % contain alkali metal ion. Metal ion–binding proteins are extensively involved in many different biochemical reactions, and metal ions are crucial to their functions. Identifying metal ion–binding sites is a key to understanding the biological function mechanisms of metal ion–binding proteins.
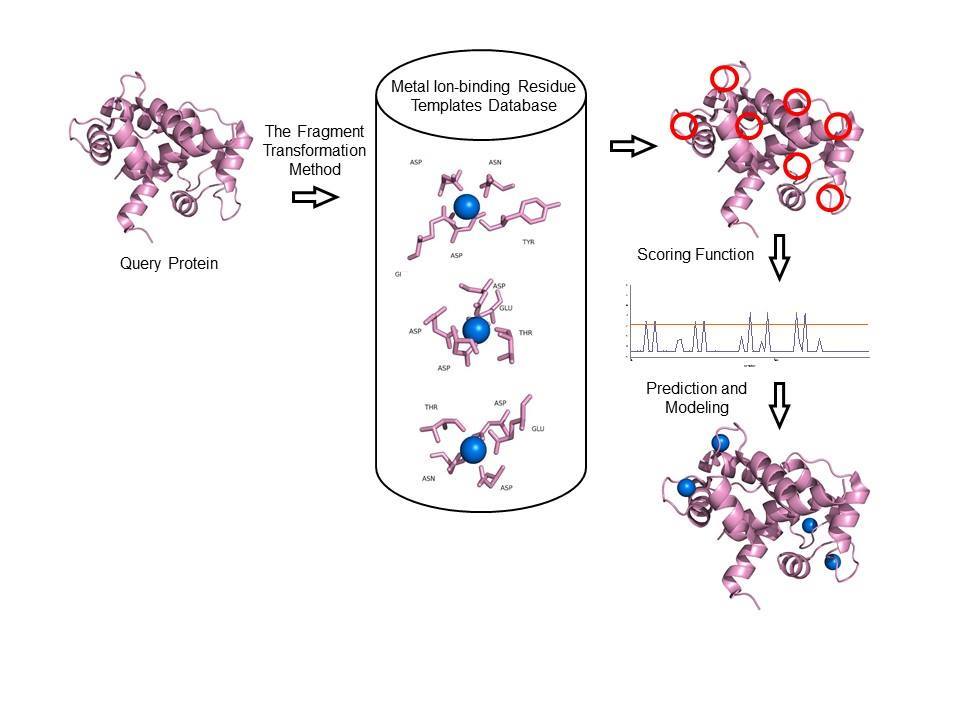
In the method of MIB2, kinds of metal ions-binding residues had been constructed, and then gathered as metal ion-binding residue templates, which are metal ion-binding residues spherical within 3.5Å of the center of the metal ion. By taking advantage of the fragment transformation method, the local structure comparison between metal ions-binding residue templates and a query protein structure with unknown binding sites was carrying out. After comparison, each residue of the query protein is assigned an alignment score using a scoring function composed of two criteria which are used to evaluate how well the sequence and structure are conserved. By setting a threshold for alignment scores, residues scoring above the threshold are predicted to be metal-binding residues. MIB2 combine both structural and sequence information to identify the local structure of protein-metal interaction sites.